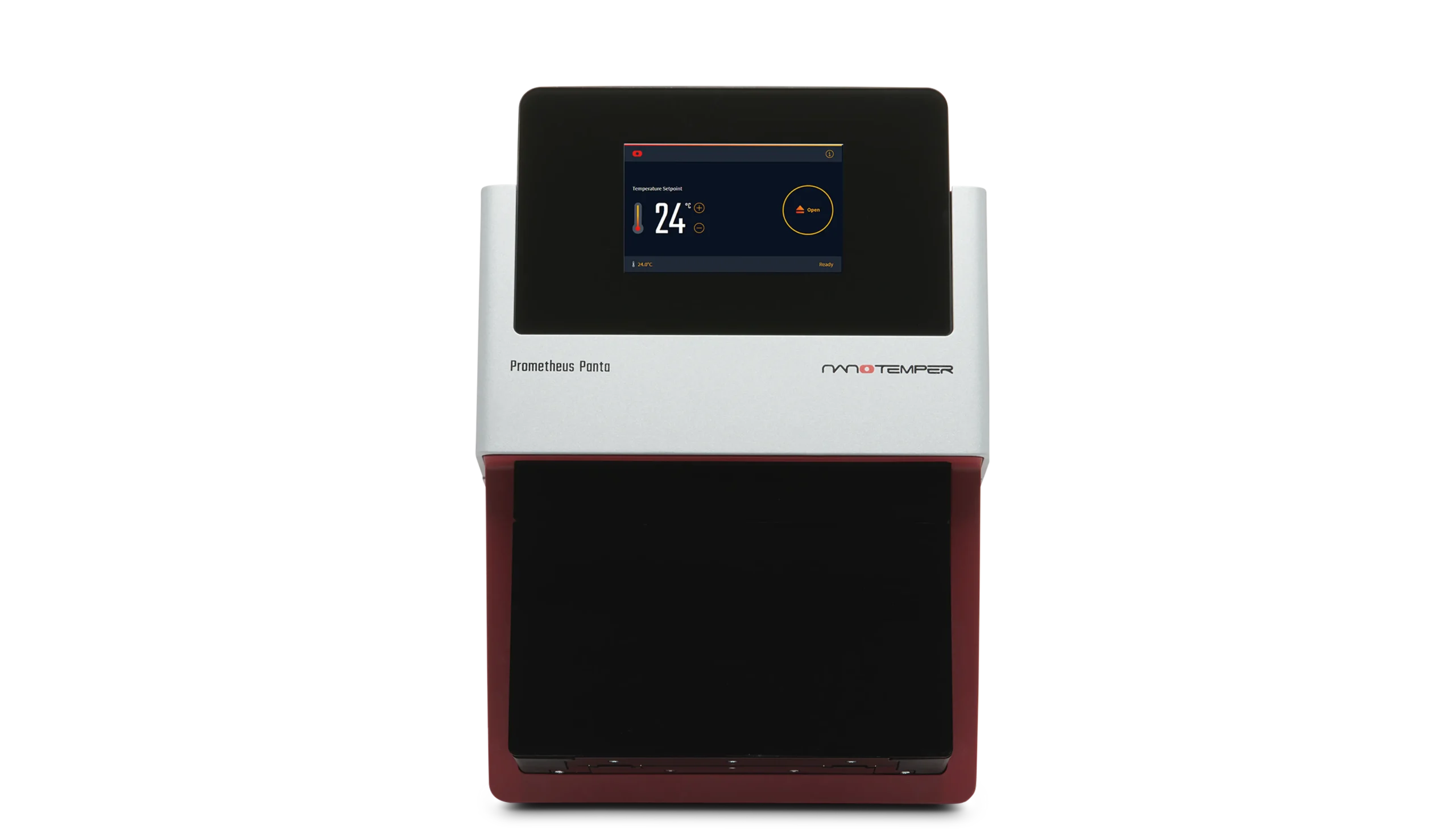

nanoDSF - nano Differential Scanning Fluorimetry.
nanoDSF is a label-free technique that monitors intrinsic protein fluorescence to measure conformational stability.
When to use nanoDSF.
nanoDSF measures conformational stability, or how well a protein maintains its folded structure under stress. The method tracks how proteins respond to thermal or chemical stress by detecting shifts in tryptophan and tyrosine fluorescence emission.
This provides quantitative stability parameters without requiring external labels or dyes. nanoDSF is critical to gain a complete stability profile for your protein.
nanoDSF is ideal when you need to:




When to use nanoDSF.
nanoDSF measures conformational stability, or how well a protein maintains its folded structure under stress. The method tracks how proteins respond to thermal or chemical stress by detecting shifts in tryptophan and tyrosine fluorescence emission.
This provides quantitative stability parameters without requiring external labels or dyes. nanoDSF is critical to gain a complete stability profile for your protein.
nanoDSF is ideal when you need to:
- Compare conformational stability across protein variants, buffer conditions, or formulations
- Rank biologics candidates based on thermal or chemical stability
- Detect subtle stability changes that other methods might miss
- Work label-free to avoid artifacts from fluorescent tags
- Screen conditions rapidly with minimal sample consumption
The science behind nanoDSF.
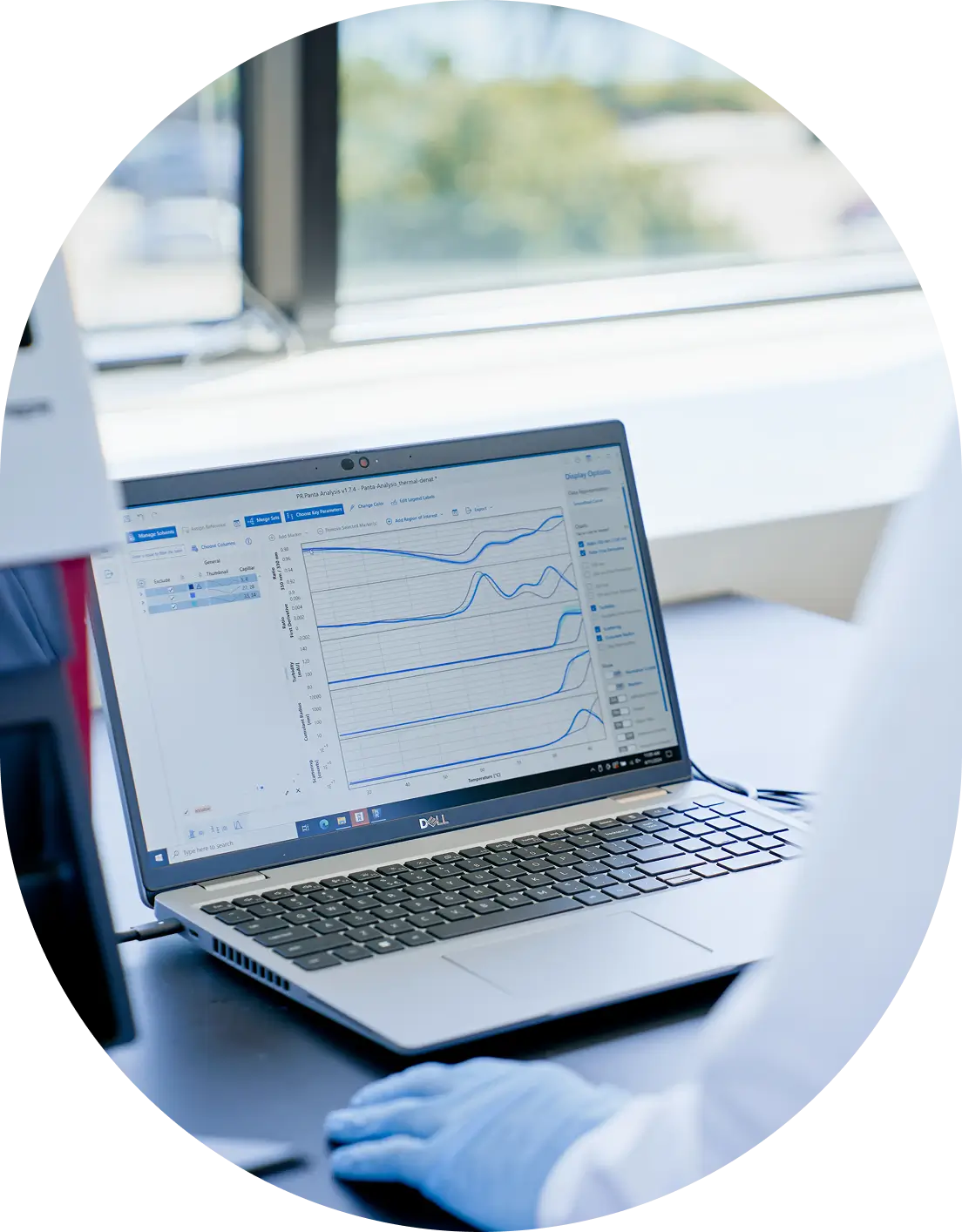

Protein folding creates distinct fluorescence environments
Proteins fold into three-dimensional structures that bury hydrophobic residues, including tryptophan and tyrosine, in their core. These aromatic amino acids absorb UV light at 280 nm and emit fluorescence, but the wavelength of that emission depends on their chemical environment.
When tryptophan is buried in a folded protein’s hydrophobic core, shielded from water, it emits fluorescence with a maximum around 330 nm. When the protein unfolds and tryptophan becomes exposed to the aqueous environment, the emission maximum shifts to around 350 nm. Tyrosine contributes similarly, though its shift is less pronounced.
This environment-dependent fluorescence shift is the foundation of nanoDSF. By monitoring the ratio of emission at 350 nm to 330 nm (F350/F330), you can track the folding state of your protein in real time.
Thermal and chemical stress trigger unfolding
During a nanoDSF experiment, proteins are subjected to a controlled stress gradient—either increasing temperature or increasing concentration of a chemical denaturant like urea or guanidinium chloride.
As stress increases, proteins begin to unfold. The population shifts from predominantly folded (low F350/F330 ratio) to predominantly unfolded (high F350/F330 ratio). Plotting this ratio against temperature or denaturant concentration produces a sigmoidal transition curve.
The inflection point of this curve—where the slope is steepest—represents the midpoint of the unfolding transition. This is where 50% of the protein population is folded and 50% is unfolded.
First derivative analysis sharpens the view
Taking the first derivative of the F350/F330 ratio with respect to temperature (or denaturant concentration) converts the sigmoidal curve into a peak. The maximum of this peak corresponds precisely to the inflection point, making it easier to identify the transition midpoint and compare stability across samples.
Parameters you measure with nanoDSF.
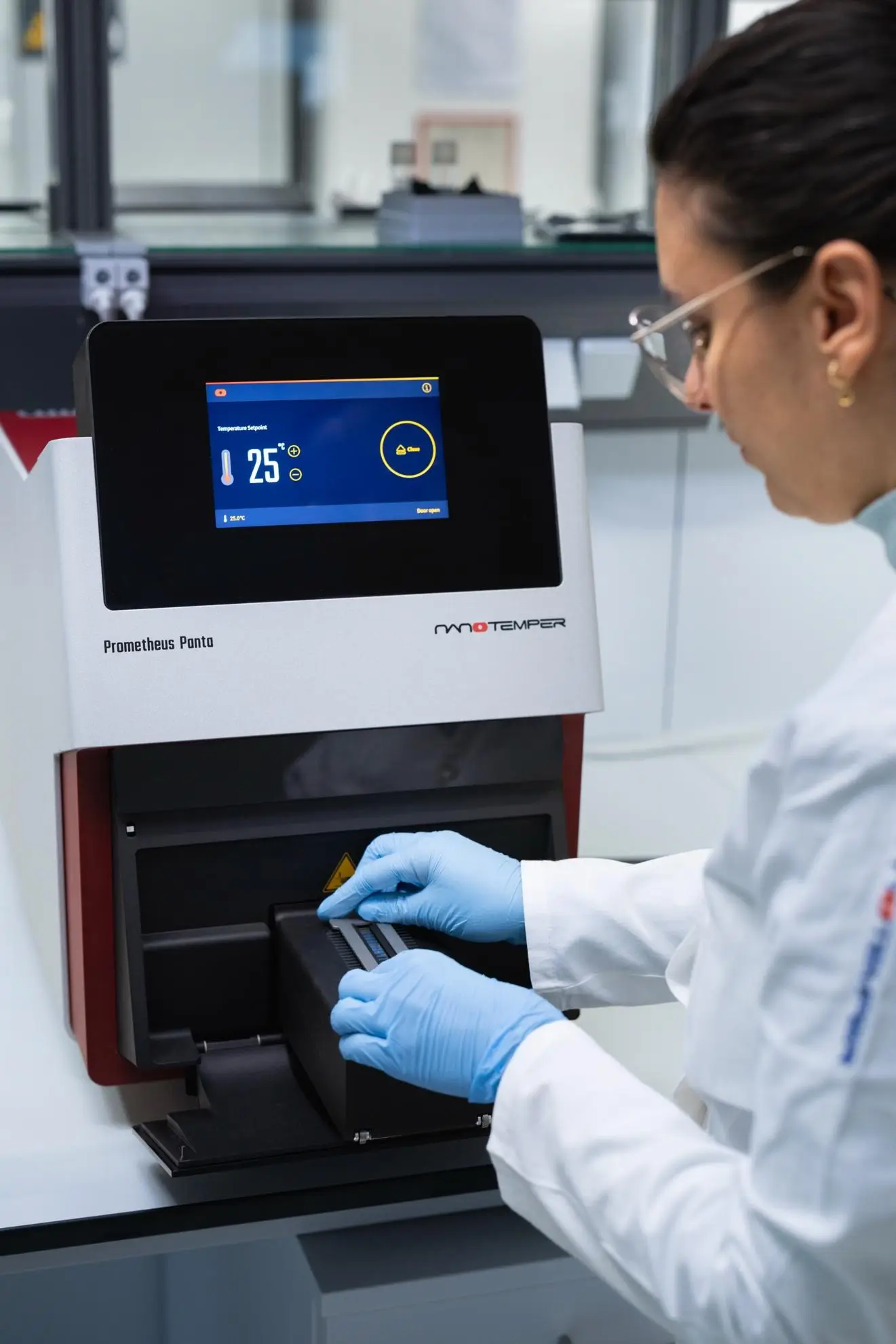





Parameters you measure with nanoDSF.

Melting temperature (Tm)
The temperature at which 50% of the protein is folded and 50% is unfolded. Higher Tm indicates greater thermal stability. Tm is the most commonly reported stability parameter and is useful for ranking candidates or conditions.

Onset of unfolding (Tonset)
The temperature where the protein first begins to unfold, identified as the point where the fluorescence ratio starts to deviate from baseline. Some mutations or buffer changes shift Tonset without significantly affecting Tm, making this parameter valuable for detecting subtle stability differences.

Transition slope (sharpness)
The steepness of the unfolding transition, measured as the slope at the inflection point. A steep slope indicates a highly cooperative unfolding process: the protein stays tightly folded until a critical temperature, then unfolds rapidly. A shallow slope suggests a less cooperative, more gradual unfolding.

Chemical denaturation midpoint (Cm or C50)
The concentration of denaturant at which 50% of the protein is unfolded. Like Tm, higher Cm values indicate greater stability.

Free energy of unfolding (ΔG)
The thermodynamic stability of the protein at a given temperature, representing the energy difference between folded and unfolded states. More positive ΔG means the protein is more stable. ΔG compares stability changes between conditions (e.g., wild-type vs. mutant).
Better show than tell. See how DLS generates information about your sample.
How nanoDSF complements other techniques.
nanoDSF measures conformational stability—how well a protein maintains its folded structure under stress. This is distinct from colloidal stability, which describes how well proteins stay in solution without aggregating.
A protein can be conformationally stable (high Tm) but colloidally unstable (prone to aggregation). Conversely, a protein might unfold easily (low Tm) but remain soluble. For a complete stability profile, combine nanoDSF with techniques like DLS, SLS, or backreflection.
In this graph, the protein is highly soluble and does not show signs of aggregation. This indicates that its formulation is optimal. However, based on the nanoDSF data, we see that the protein is not thermally stable under these conditions. Relying on DLS only could lead to development of the wrong candidate.
“NanoTemper helps us turn challenging biophysical tasks into routine workflows. Their intuitive solutions give us reliable data faster, so our teams can focus on advancing drug candidates.”
“NanoTemper helps us turn challenging biophysical tasks into routine workflows. Their intuitive solutions give us reliable data faster, so our teams can focus on advancing drug candidates.”
“NanoTemper helps us turn challenging biophysical tasks into routine workflows. Their intuitive solutions give us reliable data faster, so our teams can focus on advancing drug candidates.”
“NanoTemper helps us turn challenging biophysical tasks into routine workflows. Their intuitive solutions give us reliable data faster, so our teams can focus on advancing drug candidates.”
Our biophysical tools
that use nanoDSF.
Join the Newsletter
Get email updates. Your edge in biophysics. Subscribe today.
Our latest research with nanoDSF.
Frequently Asked Questions
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Integer tellus tellus, ornare a felis quis, semper tempus elit. Donec in facilisis magna.
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Integer tellus tellus, ornare a felis quis, semper tempus elit. Donec in facilisis magna.
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Integer tellus tellus, ornare a felis quis, semper tempus elit. Donec in facilisis magna.
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Integer tellus tellus, ornare a felis quis, semper tempus elit. Donec in facilisis magna.
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Integer tellus tellus, ornare a felis quis, semper tempus elit. Donec in facilisis magna.



